Introduction
UV-Vis spectrophotometry remains a cornerstone in biological research, offering unparalleled capabilities in quantifying biomolecules, monitoring dynamic processes, and ensuring sample purity. This article synthesizes recent advancements, emerging technologies, and practical applications of UV-Vis spectrophotometry, with a focus on its integration into modern biological workflows.
Principles of UV-Vis Spectrophotometry
Electronic Transitions and Molecular Interactions
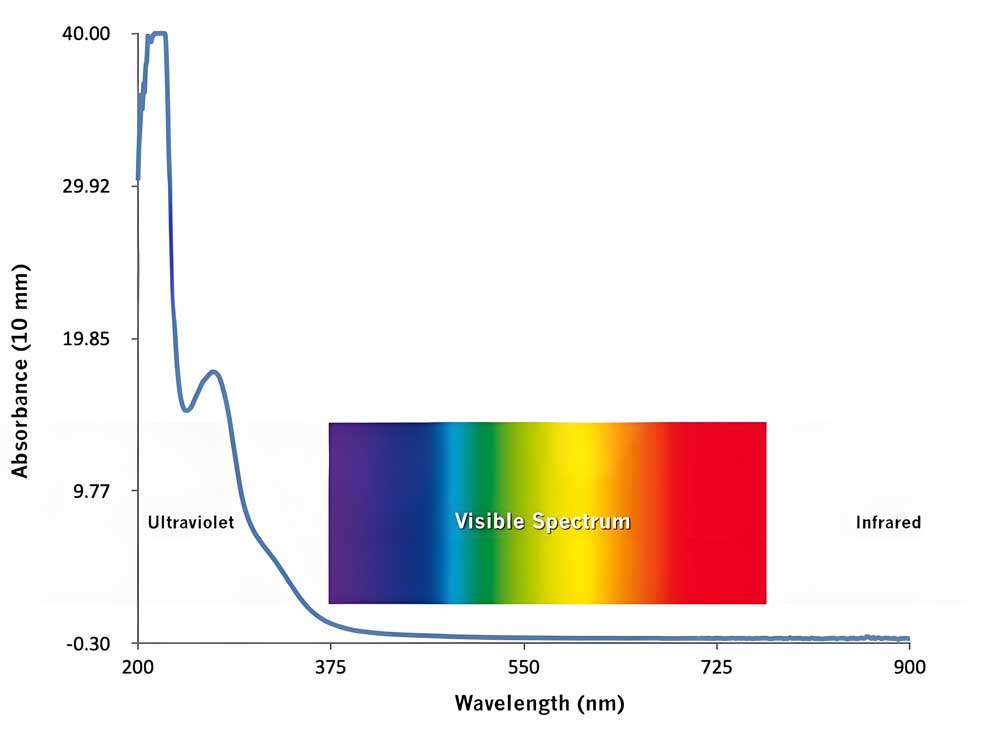
UV-Vis absorption arises from electronic transitions in biomolecules, including π→π* (aromatic systems), n→π* (heteroatoms like oxygen or nitrogen), and σ→σ* (saturated bonds). Chromophores such as tryptophan in proteins or nucleic acid bases in DNA dictate characteristic absorbance peaks (e.g., DNA at 260 nm). Deviations from the Beer-Lambert law occur in biological media due to light scattering, turbidity, or molecular aggregation, necessitating corrections for accurate quantification.
Instrumentation and Detection Systems
Modern UV-Vis spectrophotometers utilize advanced detectors (photodiode arrays, photomultiplier tubes) and light sources (deuterium lamps for UV, tungsten for visible range). Double-beam systems enhance stability for long-term kinetic studies, while array-based detectors enable high-throughput screening of multi-analyte samples. Portable devices now support field-based applications, such as environmental monitoring or point-of-care diagnostics.
Methodological Advancements
Sample Preparation and Error Mitigation
- Turbidity Management: Centrifugation or filtration reduces scattering in cell lysates or bacterial cultures.
- Solvent Optimization: Buffers like Tris-HCl maintain biomolecule stability, while avoiding solvents with UV absorption (e.g., some organic solvents).
- Calibration Standards: Traceable standards (e.g., NIST-certified BSA for proteins) ensure reproducibility across laboratories.
AI-Driven Spectral Analysis
Machine learning models, such as UV-adVISor, predict UV-Vis spectra from molecular structures using attention-based neural networks. These tools accelerate drug discovery by identifying reaction products or impurities in complex mixtures, achieving high accuracy (RMSE <0.065, R² >0.7).
Applications in Biological Research
Quantitative Biomolecule Analysis
- Nucleic Acids: Concentration and purity assessment via A260/A280 ratios, with deviations indicating protein contamination.
- Proteins: Direct quantification at 280 nm (aromatic residues) or via colorimetric assays (Bradford, BCA). Hyperspectral imaging enhances spatial resolution in tissue samples.
Enzyme Kinetics and Metabolite Monitoring
Real-time tracking of NADH/NADPH at 340 nm enables analysis of dehydrogenase activity. Time-resolved spectrophotometry captures transient intermediates in enzymatic cycles, while mid-UV spectroscopy (200–300 nm) quantifies peptides and amino acids without interference from water vibrations.
Quality Control and Impurity Profiling
UV-Vis ensures batch consistency in biopharmaceuticals by detecting contaminants (e.g., host cell proteins) or verifying drug stability. Advanced algorithms deconvolute overlapping spectra in multi-component systems, such as cell culture media.
Instrumentation Innovations
| Type | Advantages | Biological Use Cases |
|---|---|---|
| Double-Beam | Reduced baseline drift, ideal for kinetic assays | Enzyme activity monitoring, long-term cell culture studies |
| Array-Detector | Full-spectrum scans in seconds, multiplexed detection | High-throughput drug screening, multi-omics integration |
| Portable | Battery-operated, field-deployable | Environmental sampling, on-site diagnostics |
Challenges and Solutions
Beer-Lambert Law Deviations
Biological matrices (blood, saliva) often violate linearity due to scattering or fluorescence. Integrating sphere attachments or Mie theory corrections improve accuracy in turbid samples.
Standardization and Reproducibility
Adherence to ISO 17025 protocols and use of certified reference materials minimize inter-lab variability. Automated systems reduce human error in sample handling and data analysis.
Future Perspectives
- Miniaturization and Wearable Sensors: Ultra-compact devices for real-time health monitoring (e.g., glucose levels in sweat).
- Multi-Omics Integration: Coupling with mass spectrometry or NMR for systems-level biological insights.
- Quantum Dot Enhancements: Nanomaterials boost detection sensitivity for low-abundance biomarkers.
References and further readings:
1.Schmid, F. X. (2001). Biological Macromolecules: UV-Visible Spectrophotometry. Wiley Online Library. https://doi.org/10.1038/npg.els.0003142
2.Eppendorf. (2020). UV-Vis Spectrophotometry – Easy and Quick Quantification of Nucleic Acids. Eppendorf Finland. https://www.eppendorf.com/fi – en/lab – academy/life – science/cell – biology/uv – vis – spectrophotometry – easy – and – quick – quantification – of – nucleic – acids/
3.íos – Reina, R., & Azcarate, S. M. (2023). How Chemometrics Revives the UV – Vis Spectroscopy Applications as an Analytical Sensor for Spectralprint (Nontargeted) Analysis. MDPI Chemosensors. https://doi.org/10.3390/chemosensors11010008
4.Tellinghuisen, J. (2000). Statistical Error Calibration in UV – Vis Spectrophotometry. Applied Spectroscopy. https://doi.org/10.1366/0003702001949537
Conclusion
UV-Vis spectrophotometry continues to evolve, driven by technological innovations and interdisciplinary applications. By addressing challenges such as matrix effects and leveraging AI-driven tools, it remains indispensable in advancing biomedical research, environmental science, and industrial quality control.
Leo Bios
Hello, I’m Leo Bios. As an assistant lecturer, I teach cellular and
molecular biology to undergraduates at a regional US Midwest university. I started as a research tech in
a biotech startup over a decade ago, working on molecular diagnostic tools. This practical experience
fuels my teaching and writing, keeping me engaged in biology’s evolution.







Leave a Comment
Your email address will not be published. Required fields are marked *